VariantSpark: Machine learning for ultra-high dimensional data
Machine learning analysis suite specifically tailored for clinical and genomic data analysis.
The challenge
Clinical and genomic datasets are becoming more complex.
Researchers are sourcing data from more people and collecting more clinical information per person, resulting in high-dimensional data.
Current machine learning (ML) solutions are not designed to deal with ultra high-dimensional data. A common work-around has been to pre-filter the “raw” datasets or limit the analysis to independent markers only, in order to reduce the computational cost.
However, eliminating information or only assessing independent drivers in complex diseases can potentially bias results and limit the accuracy.
Our response
We developed VariantSpark, a machine learning framework able to deal with the depth and complexity of clinical data.
VariantSpark can cluster clinical profiles or identify disease-associated features from trillions of data points (thousands of samples with millions of features each) in just 30 minutes.
VariantSpark is available through the Digital Marketplace on Amazon Web Services (AWS). It can be deployed on Microsoft Azure and Google Cloud Platform (GCP, via Terra notebook).
The underlying random forest algorithms offer explainable machine learning. Which features contributed in what proportion to the overall prediction outcome are visualised in an interactive graph.
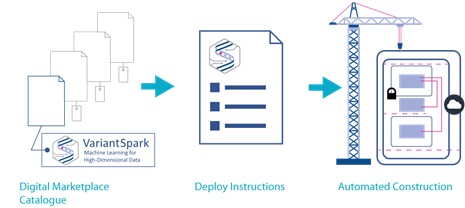
Obtaining, deploying and constructing (automatic using cloud construction) VariantSpark.
Benefits
VariantSpark is 90% faster than traditional compute frameworks and requires 80% fewer samples to detect statistically significant signal.
It detects stronger predictive markers by considering the full dataset and the interaction between multiple clinical features.
In a genome-wide-association case/control set-set up, VariantSpark can identify higher order interactions between genomics features.
In the clinical setting, VariantSpark enables personalised genomic insights at point-of-care, by finding patients-like-mine based on genomic similarity to other patients.

VariantSpark is machine learning for high-dimensional data. It can be used for applications such as disease gene discovery.
The Australian e-Health Research Centre (AEHRC) is CSIRO's digital health research program and a joint venture between CSIRO and the Queensland Government. The AEHRC works with state and federal health agencies, clinical research groups and health businesses around Australia.

